Discovery of AH molecular marker by proteomic evaluation
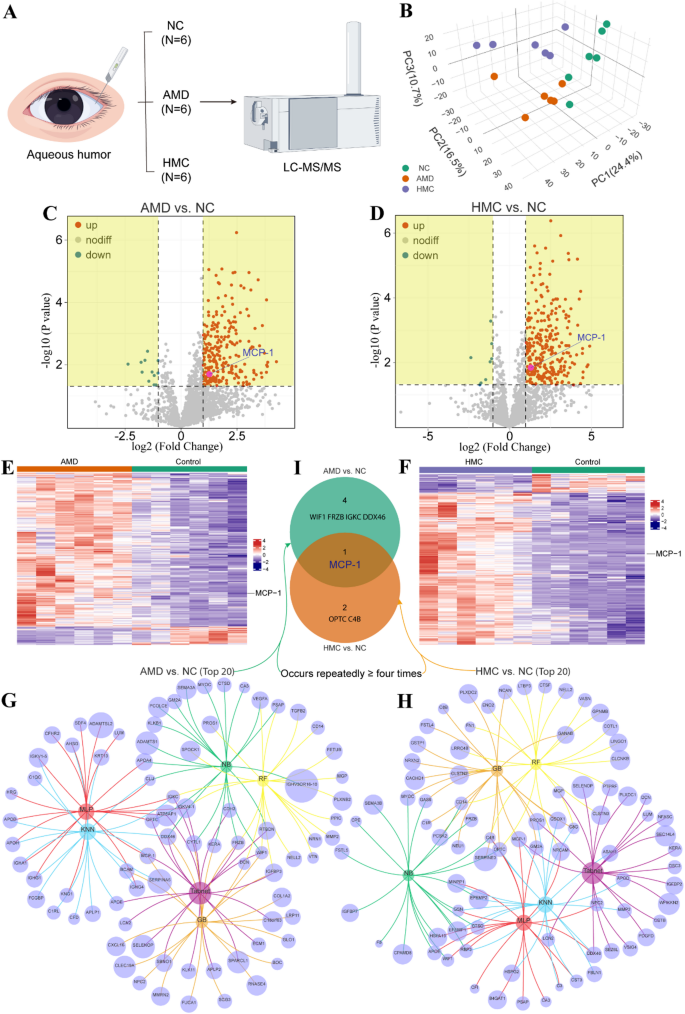
To analyze novel doable molecular markers for widespread screening and early discovery of blinding eye ailments, a complete proteomic evaluation was carried out on a complete of six AH samples sourced from ARC, which served as regular controls (NC), together with one other six AH samples from AMD and 6 AH samples from HMC sufferers (Fig. 1A). Utilizing in-depth proteome profiling, we recognized 2,182 expressed proteins in a complete of 18 AH samples from three teams. Principal Element Evaluation (PCA) confirmed clear distinctions between these three teams (Fig. 1B). Volcano plots demonstrated molecular markers with vital variations, outlined as Bonferroni-adjusted P-value < 0.05 and log2 FC absolute worth > 1, between AMD and NC (Fig. 1C) and HMC and NC (Fig. 1D). Moreover, a heatmap illustrated the considerably upregulated and downregulated markers within the AMD vs. NC comparability (Fig. 1E) and the HMC vs. NC comparability (Fig. 1F). The highest 20 most essential markers had been ranked primarily based on SHAP values derived from every machine studying algorithm (Determine S2). Within the AMD vs. NC evaluation, MCP-1, WIF1, FRZB, IGKC, and DDX46 constantly appeared within the high 20 rankings throughout not less than 4 algorithms (Fig. 1G). Within the HMC vs. NC evaluation, MCP-1, OPTC, and C4B had been constantly recognized within the high 20 rankings throughout not less than 4 algorithms (Fig. 1H). Lastly, MCP-1 was recognized as a typical marker, rising from the intersection of the NC vs. AMD and NC vs. HMC analyses (Fig. 1I). Due to this fact, by particularly recognizing and sensitively monitoring MCP-1, it’s going to possible assist in the early detection of the blinding eye ailments AMD and HMC.
Screening and identification of markers for blinding eye ailments. (A) Schematic diagram of proteomics evaluation of human AH samples. (B) PCA rating plot exhibiting clear separation among the many AMD, HMC, and NC teams, reflecting distinct protein expression profiles. (C) Volcano plot of differential proteins between the AMD and NC teams, and (D) the HMC and NC teams, with considerably altered proteins outlined by a Bonferroni-adjusted P-value < 0.05 and|Log2 Fold Change| >1. (E) Heatmap depicting the expression patterns of considerably upregulated and downregulated proteins within the AMD vs. NC, and (F) HMC vs. NC comparability. (G) High 20 most essential proteins recognized primarily based on SHAP values derived from a number of machine studying algorithms within the AMD vs. NC and (H) HMC vs. NC comparability. (I) Venn diagram highlighting MCP-1 as a significant differential protein recognized as a candidate marker in each comparisons (AMD vs. NC and HMC vs. NC)
Screening and building of MCP-1-targeting aptamers
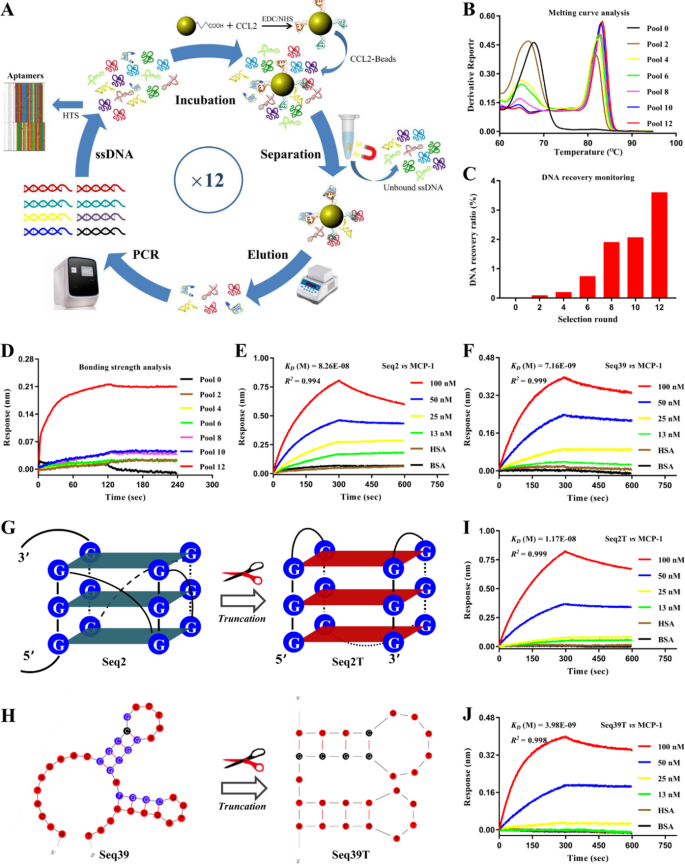
To acquire extremely delicate BRMs concentrating on MCP-1, aptamers had been screened utilizing the SELEX approach (Fig. 2A). Throughout in vitro screening, we designed a SELEX program consisting of a optimistic choice step to retain ssDNA that particularly bind to MCP-1, together with a unfavourable choice step to remove ssDNA that nonspecifically bind to bead carriers. Moreover, SELEX situations had been custom-made to permit the MCP-1 binding sequence to compete with free counter-targets, leading to aptamers with excessive recognition exercise. Nonetheless, the enrichment of nonspecific ssDNA was unavoidable; thus, SELEX development was monitored to make sure profitable aptamer acquisition. We carried out a complete evaluation of the melting curves, recoveries, and binding strengths of the screened libraries to find out aptamer convergence. As proven in Fig. 2B, the melting peak of the libraries shifted from 81.4 °C in Pool 2 to 83.6 °C in Pool 12, reaching a plateau. The restoration share additionally confirmed a big improve in Pool 12 (Fig. 2C). Moreover, Pool 12 started to exhibit an MCP-1 binding sign depth stronger than that of Pool 0 (Fig. 2D). Based mostly on these outcomes, we stopped the choice course of and carried out high-throughput sequencing of the enriched Pool 12 library, together with a number of sequence comparability evaluation utilizing Clustal X software program. Sequences had been grouped into two households primarily based on homology (Determine S3). The chosen consultant aptamers, Seq2 and Seq39, exhibited good affinity and specificity for MCP-1; their affinity constants had been 82.6 nM and seven.16 nM, respectively (Fig. 2E and F).
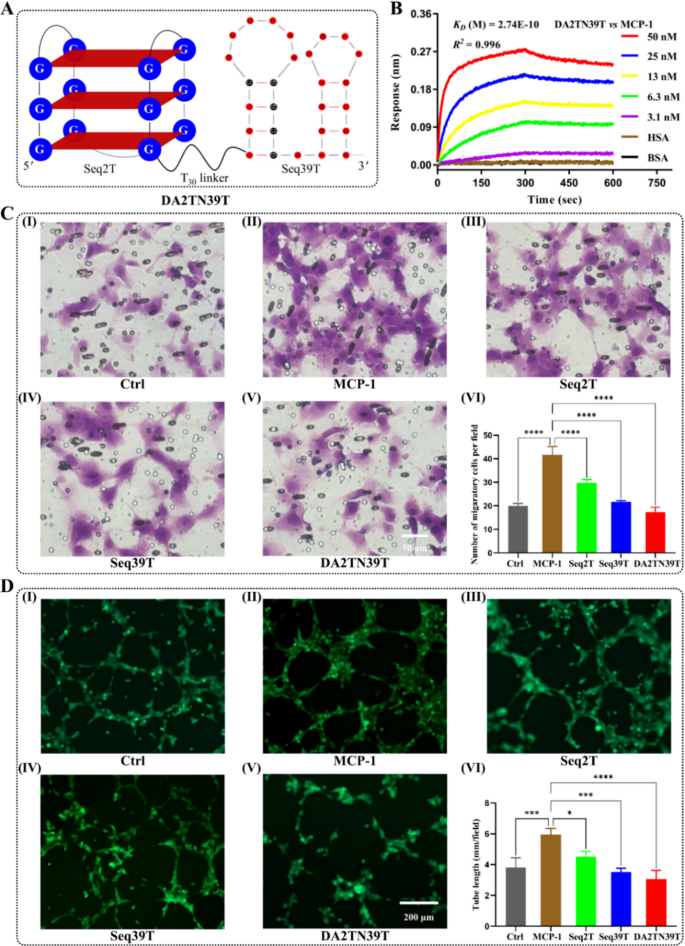
After screening, we optimized and modified the preliminary aptamers. QGRS and Mfold net server predictions revealed that Seq2 folded right into a G-quadruplex (Fig. 2G), whereas Seq39 had a double stem-loop construction (Fig. 2H). Truncation of redundant sequences at each ends of the aptamer buildings yielded the core buildings Seq2T and Seq39T, which retained excessive affinity and specificity for MCP-1 (Fig. 2I and J). To reinforce aptamer sensitivity to be used as BRMs, we utilized a bivalent engineering technique that concerned two linking monomers, thereby producing dimeric aptamers [32,33,34]. Particularly, we linked Seq2T and Seq39T with a polythymidine (T30) linker, yielding two homodimeric aptamers (DA2TN2T and DA39TN39T) and one heterodimeric aptamer (DA2TN39T), the place DA signifies dimeric aptamers, 2 T signifies Seq2T, N signifies and, in addition to 39 T signifies Seq39T (Desk S1). BLI detection revealed that the OkayD values of homodimeric aptamers DA2TN2T and DA39TN39T for MCP-1 had been 34.9 nM and eight.05 nM, respectively (Determine S4 and S5). This means that the aptamer monomer binds to just one epitope on MCP-1, whereas the formation of homodimers barely reduces the interplay between them. In distinction, the heterodimeric DA2TN39T confirmed a considerably enhanced binding capability relative to its monomeric counterparts, with a OkayD worth lowered to 0.274 nM (Fig. 3A and B), which can be as a result of totally different binding epitopes of Seq2T and Seq39T on MCP-1. To additional validate the considerably elevated concentrating on efficiency of DA2TN39T, we carried out a organic perform inhibition assay involving MCP-1. As proven in Fig. 3C and D, we investigated the consequences of MCP-1-targeting aptamers on the migration and angiogenesis of RF/6A cells. The cells had been untreated or pretreated with exogenous MCP-1 protein. Moreover the exogenous MCP-1, the aptamers Seq2T, Seq39T, and DA2TN39T had been pre-added within the experimental teams. The outcomes confirmed that Seq2T, Seq39T, and DA2TN39T all considerably inhibited the migration of RF/6A cells (Fig. 3C, VI); the dimeric aptamer DA2TN39T exhibited the best impact. The tube formation assay additionally revealed that DA2TN39T had a superior rescue impact on the MCP-1-induced improve in angiogenesis potential, in contrast with Seq2T or Seq39T (Fig. 3D, VI). Due to this fact, we obtained a extremely delicate bivalent aptamer, DA2TN39T. As a novel BRM that achieves excessive binding affinity, goal specificity and structural stability by concurrently interacting with two epitopes on MCP-1.
MCP-1 aptamer screening and optimization. (A) Schematic diagram of magnetic bead–primarily based SELEX. (B) Melting curve evaluation of enriched screening libraries utilizing the QuantStudio™ Actual-Time PCR System. (C) Monitoring of ssDNA restoration from screening libraries through Qubit® fluorometry. (D) Biolayer interferometry evaluation of binding energy within the enriched screening libraries. (E, F) Biolayer interferometry evaluation of intermolecular interactions of the unique aptamers Seq2 and Seq39 with MCP-1. (G, H) Schematic diagram of the folded buildings of Seq2 and Seq39 earlier than and after truncation optimization. (I, J) Biolayer interferometry evaluation of the affinity and specificity of the truncated aptamers Seq2T and Seq39T for MCP-1
Dimeric aptamer building and identification. (A) Schematic diagram of the constructed heterodimeric aptamer DA2TN39T. (B) Biolayer interferometry evaluation of the affinity and specificity of DA2TN39T for MCP-1. (C) RF/6A cells had been untreated (Ctrl; I) or pretreated with MCP-1 (II) with or with out the Seq2T (III), Seq39T (IV), or DA2TN39T (V) for twenty-four h. Transwell assays had been then carried out to evaluate the migration of RF/6A cells (n = 3; VI). (D) RF/6A cells had been untreated (Ctrl; I) or pretreated with MCP-1 (II) with or with out the aptamers Seq2T (III), Seq39T (IV), or DA2TN39T (V) for twenty-four h. Tube formation potential was detected and analyzed (n = 3; VI). *p (<) 0.05, ***p (<) 0.0005, and ****p (<) 0.0001 in contrast with MCP-1 therapy
Recognition mechanism of MCP-1-targeting dimeric aptamer DA2TN39T
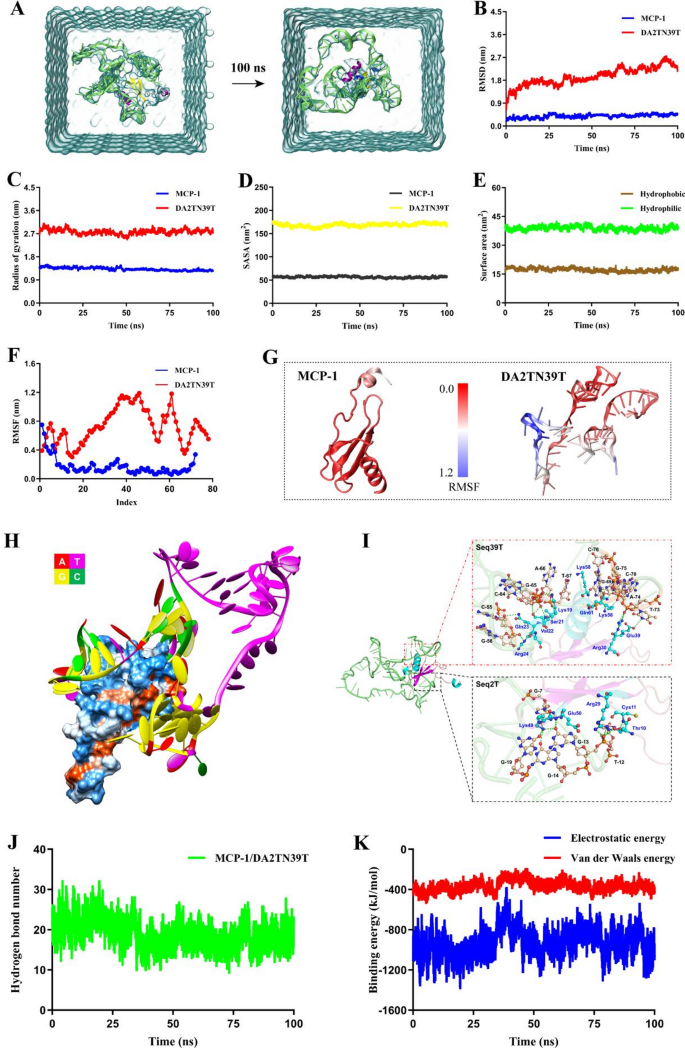
MD simulations had been used to discover the molecular mechanism by which the DA2TN39T (Determine S6) targets MCP-1 (Determine S7). As proven in Fig. 4A, the construction of the DA2TN39T-MCP-1 complicated didn’t considerably change earlier than and after MD simulations, indicating that MCP-1 kinds a steady complicated after binding with DA2TN39T. The basis imply sq. deviation (RMSD) of MCP-1 throughout the system remained comparatively steady throughout MD simulations (Fig. 4B). Though the RMSD worth of DA2TN39T partially fluctuated in the course of the simulations, this may occasionally have occurred as a result of the linker portion of the dimeric aptamer was free in resolution. Total, the imply RMSD values of MCP-1 and DA2TN39T had been 0.389 ± 0.067 nm and 1.952 ± 0.348 nm, respectively. Moreover, Fig. 4C signifies that in MD simulations, the Rg values of MCP-1 and DA2TN39T had been comparatively steady, with imply Rg values of 1.326 ± 0.056 nm and a pair of.776 ± 0.094 nm, respectively. These outcomes advised that MCP-1 and DA2TN39T can type a extremely steady complicated. Subsequently, we analyzed adjustments within the structural properties of MCP-1 and DA2TN39T throughout MD simulations. As proven in Fig. 4D, the solvent-accessible floor space (SASA) worth of MCP-1 was comparatively steady throughout MD simulations, with a imply worth of 56.377 ± 1.652 nm2. Though the SASA worth of DA2TN39T barely decreased originally of the simulations, it later stabilized close to 50 ns with a imply worth of 169.874 ± 2.527 nm2. To additional characterize structural alterations in MCP-1 upon binding to DA2TN39T, we analyzed adjustments within the protein’s hydrophobic and hydrophilic floor areas. As proven in Fig. 4E, the hydrophobic and hydrophilic floor areas of the protein remained comparatively steady throughout MD simulations, with imply values of 38.839 ± 1.072 nm2 and 17.207 ± 0.868 nm2, respectively, indicating that the construction of MCP-1 doesn’t considerably change upon binding to DA2TN39T. Moreover, Fig. 4F and G presents the pliability distributions of amino acids in MCP-1 and bases in DA2TN39T throughout the system. Along with the better flexibility of the N-terminal amino acid residues in MCP-1, the general root imply sq. fluctuation (RMSF) of the amino acid residues was low. This outcome could also be as a result of stabilizing impact of aptamer binding on the protein construction. The buildings at each ends of DA2TN39T had low flexibility due to their direct binding to MCP-1. Nonetheless, the unbound linker area could also be strongly influenced by solvent interactions, leading to massive base actions. Due to this fact, upon protein binding, the binding area of the aptamer additionally shows enhanced structural stability.
To additional elucidate the important thing binding websites and driving forces underlying the interactions between the DA2TN39T and MCP-1, we investigated their binding mode. Determine 4H reveals that the Seq2T and Seq39T areas at each ends of the dimeric aptamer had been primarily sure to the hydrophilic area of MCP-1, the place they fashioned hydrogen bonds. The central T30 constituted a linker between the 2 finish areas, functioning as a extremely steady MCP-1 “clamp.” As demonstrated in Fig. 4I, the primary bases in Seq39T that contributed to hydrogen bond formation had been C-55, G-56, C-64, G-65, A-66, T-67, G-69, C-70, T-73, A-74, G-75, and C-76. The amino acids in MCP-1 that sure to this area included Lys19, Ser21, Val22, Gln23, Arg24, Arg30, Glu39, Lys56, Lys58, and Gln61. The primary bases in Seq2T that participated in hydrogen bond formation had been G-7, T-12, G-13, G-14, and G-19. The amino acids that sure to this area in MCP-1 included Thr10, Cys11, Arg29, Lys49, and Glu50. Total, DA2TN39T kinds an intensive community of hydrogen bonds with MCP-1. As a result of hydrogen bonds are essential elements of aptamer–protein recognition, we analyzed the variety of hydrogen bonds between DA2TN39T and MCP-1. As proven in Fig. 4J, hydrogen bond interactions between DA2TN39T and MCP-1 considerably fluctuated throughout the first 40 ns; nevertheless, it stabilized from 50 to 100 ns, with a imply of 19.01 hydrogen bonds. Moreover, the binding power between DA2TN39T and MCP-1 remained comparatively steady throughout MD simulations (Fig. 4Okay), with imply electrostatic and van der Waals energies of −922.602 ± 117.589 kJ/mol and − 366.083 ± 38.092 kJ/mol, respectively, from 50 to 100 ns. These findings indicated that sturdy intermolecular interactions had been current, enabling steady complicated formation. Total, our outcomes clarified the mechanism by which the dimeric aptamer DA2TN39T acknowledges targets; they supply a powerful theoretical foundation to assist its improvement and software as a BRM for MCP-1.
Recognition mechanism evaluation. (A) Conformation of the DA2TN39T–MCP-1 complicated earlier than and after MD simulations. Variations in RMSD (B), Rg (C), and SASA (D) of MCP-1 and DA2TN39T based on simulation time. (E) Hydrophobic and hydrophilic floor areas of the MCP-1 as a perform of MD simulation time. (F) RMSF distributions of MCP-1 and DA2TN39T (G depict flexibility distributions of the corresponding protein and aptamer buildings). (H) Patterns of DA2TN39T binding to hydrophilic and hydrophobic surfaces of MCP-1 (blue and orange areas on protein floor characterize hydrophilic and hydrophobic areas, respectively). (I) Websites of binding between aptamer bases and MCP-1 amino acid residues (inexperienced dashed traces point out hydrogen bonds). (J) Numbers of hydrogen bonds and (Okay) binding energies between DA2TN39T and MCP-1 based on MD simulation time
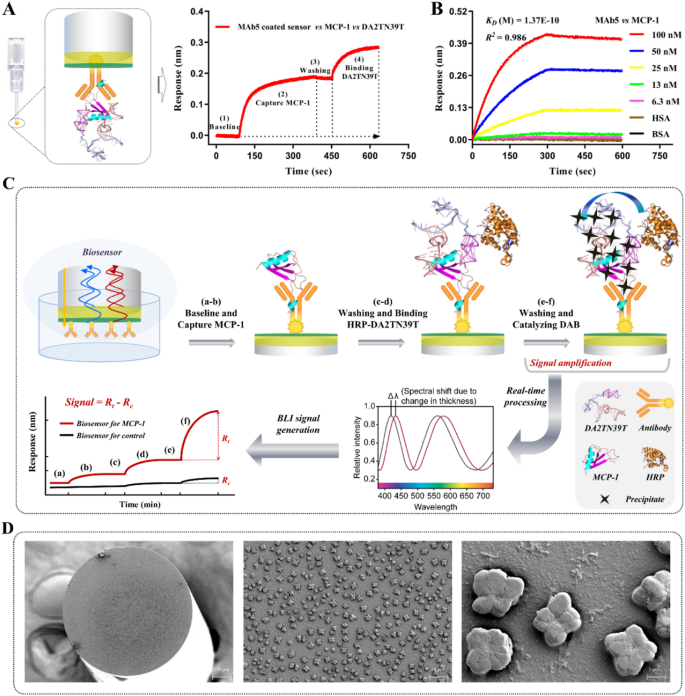
Building and precept of molecular detection know-how. (A) BLI sandwich assay–primarily based screening of antibodies that bind MCP-1 at the side of aptamers. (B) Biolayer interferometry evaluation of the binding affinity and concentrating on specificity of MAb5 for MCP-1. (C) Schematic diagram of molecular detection know-how primarily based on antibody–aptamer pairing binding and BLI sensing for MCP-1. (D) Consultant scanning electron micrograph of a biosensor with insoluble precipitates on its floor
Molecular detection know-how primarily based on antibody–aptamer focused pairing and BLI sensing for MCP-1
For prime-specificity detection of MCP-1, we deliberate a molecular binding technique of one-to-one pairing with the goal to beat the inadequate selectivity of single-material BRMs. Utilizing a BLI sandwich screening assay, monoclonal antibodies (MAb) 1 to 7 had been individually immobilized on the sensor floor, then sequentially mixed with MCP-1 and DA2TN39T. As proven in Fig. 5A, MAb5 and DA2TN39T displayed focused paired binding to MCP-1, yielding a secondary response sign. Moreover, MAb5 confirmed comparable MCP-1 binding affinity (OkayD of 0.137 nM vs. 0.274 nM; Fig. 5B) and specificity relative to DA2TN39T, guaranteeing the formation of extremely steady ternary complexes. Thus, MAb5 was immobilized on the sensor floor because the seize BRM, whereas HRP–DA2TN39T carried out pair binding as a detection BRM, considerably amplifying the sensing sign. As indicated in Fig. 5C, the MAb5-assembled sensor was equilibrated in binding buffer for 1.5 min, then incubated for five min with analyte samples to seize free MCP-1. Subsequent, the sensor was washed for 1.5 min, then incubated for a further 5 min in a nicely containing HRP-labeled aptamer (HRP–DA2TN39T) to type a ternary complicated consisting of antibody–MCP-1–aptamer. The sensor was briefly washed (1.5 min), adopted by submersion in diaminobenzidine (DAB) substrate resolution; this course of led to insoluble crystal precipitation on the sensor floor (Fig. 5D). The crystal precipitation precipitated a considerable change within the optical density of the biosensor layer and induced a big spectral shift (Δλ), amplifying the sensing sign.
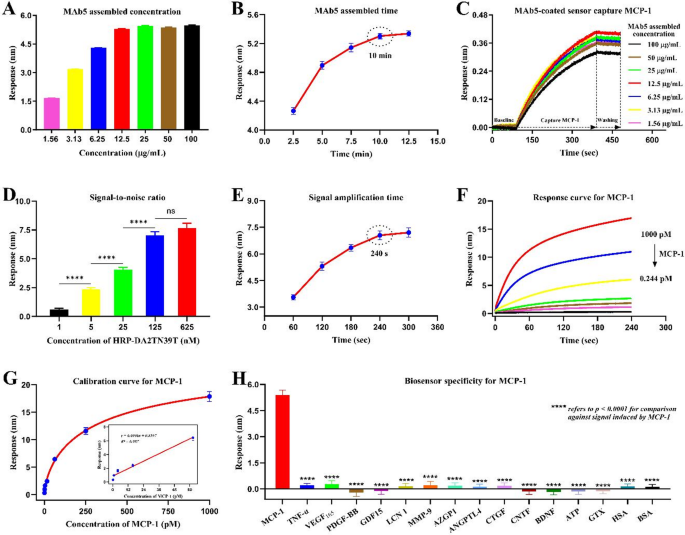
To maximise detection effectivity, we investigated the optimum meeting density and time for MAb5 on the sensor interface. As proven in Fig. 6A, the measured response elevated because the MAb5 focus approached 12.5 µg/mL, after which the sign didn’t considerably change. Determine 6B reveals the change in sensor response after incubation with 12.5 µg/mL biotin-labeled MAb5 for varied intervals. We noticed regular sign enhancement till 10 min of meeting time, when saturation was reached. At the moment, it exhibited optimum seize effectivity for the MCP-1 (Fig. 6C). Due to this fact, underneath this situation, the biosensor interface not solely achieved the utmost meeting protection of antibody MAb5, but additionally allowed the lodging of enough MCP-1 molecules in its spatial construction. Subsequent, we optimized the paired-binding BRM, HRP–DA2TN39T. Theoretically, the upper its focus, the better the variety of antibody–MCP-1–aptamer complexes fashioned. As proven in Fig. 6D, there was a transparent dose-response relationship between the HRP–DA2TN39T focus and sign stage. A wonderful signal-to-noise ratio was obtained with 125 nM of HRP–APT2TM, which signifies that not solely was there sufficient protection of the ultimate complicated on the sensing interface, but additionally the sign amplification effectivity was optimum. Moreover, the enzymatic response time in DAB resolution was important for producing enough crystal precipitates on the biosensor floor to realize increased sensitivity. Determine 6E reveals that the response considerably elevated with sign amplification time as much as 4 min, after which the response plateaued.
Optimization and software of molecular detection know-how. Optimization of MAb5 (A) meeting focus and (B) meeting time. (C) Quantity of MCP-1 captured on MAb5-coated sensors. (D) Optimization of HRP–DA2TN39T focus (****p < 0.0001; ns, no significance) and (E) sign amplification time within the detection system. (F) Time dependence of the response sign throughout the sensing system after the addition of MCP-1. The sign was recorded within the closing step of the assay. (G) Calibration curve for MCP-1, plotting response sign towards MCP-1 focus, within the vary of 0.244–1000 pM. Inset: Linear relationship between the sign and MCP-1 focus (from 0.244 to 62.5 pM). (H) Selectivity of the molecular detection know-how (****p < 0.0001 in contrast with sign induced by MCP-1)
Subsequently, underneath the above optimized experimental situations, we recorded dose-dependent adjustments within the sensing sign for MCP-1 concentrations starting from 0.244 to 1000 pM. Determine 6F reveals that the sensing system sign progressively declined because the MCP-1 focus decreased. A sigmoidal logistic four-parameter curve was obtained by plotting the sign towards the MCP-1 focus (Fig. 6G). Information factors within the calibration curve represented three impartial measurements and might be fitted to the next equation: y = (26.1904–0.3935)/[1 + (x/355.4528)^(−0.7142)] + 0.3935, R2 = 0.995. Moreover, we characterised a linear detection vary between 0.244 and 62.5 pM of MCP-1, represented by the linear regression equation: y = 0.0906x + 0.8597, R2 = 0.987. The restrict of detection (LOD) for MCP-1 was roughly 0.16 pM (S/N = 3), the place the noise stage was outlined as the usual deviation of a number of measurements from clean samples (n = 10). This sensitivity is corresponding to that of business ELISA kits. Notably, ELISA-based MCP-1 detection usually requires 4–5 h, whereas the developed molecular detection know-how using an automatic and high-throughput BLI sensing system can full the evaluation inside 30 min. As a result of selectivity is a crucial efficiency parameter in sensible purposes of molecular detection applied sciences, we carried out cross-reactivity experiments with varied potential interfering substances. Our know-how theoretically ought to exhibit good selectivity for MCP-1 as a result of the antibody–aptamer pair was used as a molecular device for one-to-one recognition and sign manufacturing. Certainly, weak responses had been noticed upon addition of different markers at equal concentrations, in sharp distinction to the sturdy sign induced by MCP-1 (Fig. 6H). Thus, our know-how detects MCP-1 in a extremely selective and delicate method with out nonspecific interactions involving different molecular markers with related buildings or coexisting. To additional consider the sensible effectiveness of the developed molecular detection know-how, we spiked human serum samples with MCP-1 and analyzed them utilizing a preset calibration curve. ELISA-based evaluation revealed that the basal stage of MCP-1 in serum was 103.67 pg/mL. Varied quantities of MCP-1 had been then added into serum and diluted 10-fold for detection. As proven in Desk S2, a great vary of restoration (96.28–105.44%) was achieved, indicating that the detection accuracy of the sensing system was not considerably altered by the serum matrix. The coefficient of variation ranged from 1.98 to five.03% (lower than 10%), indicating that the proposed molecular detection know-how has strong reproducibility and stability for evaluation of MCP-1 in precise scientific samples. Total, the developed molecular detection know-how supplies a dependable, handy, and cost-effective method for the exact identification of MCP-1, with nice potential for scientific translation.
Molecular detection know-how for diagnostic purposes in blinding eye ailments
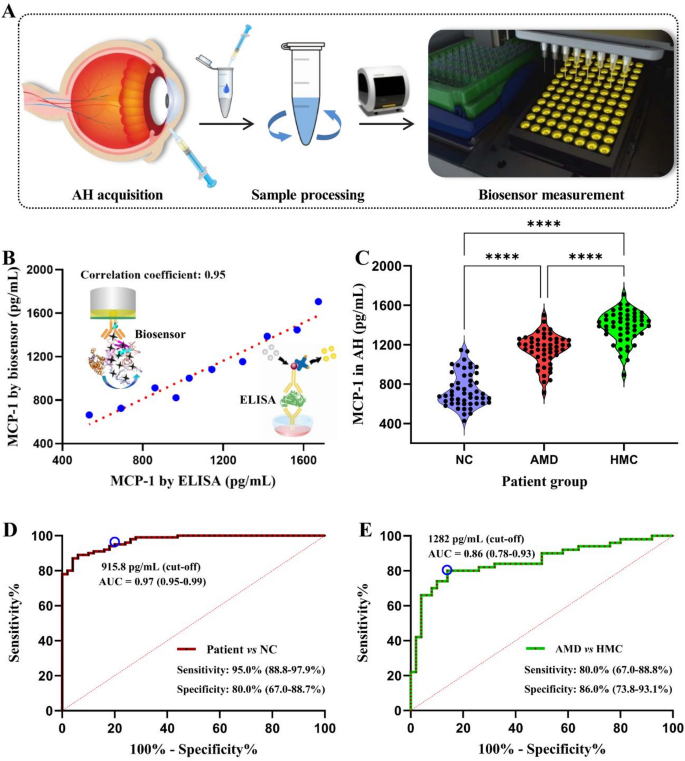
To judge the scientific diagnostic efficiency of the molecular detection know-how, we recruited 150 members, together with 50 NC, 50 AMDand 50 HMC sufferers, as a validation cohort. As proven in Fig. 7A, AH samples had been collected, processed, and subsequently analyzed utilizing our developed detection know-how or a business ELISA package. Notably, the utmost correlation coefficient between the 2 strategies for scientific AH samples was 0.95 (Fig. 7B), indicating that the concentrations detected utilizing our know-how had been corresponding to these detected utilizing the gold customary. As proven in Fig. 7C, according to the outcomes of the invention cohort, the molecular marker MCP-1 concentrations in AH samples had been considerably increased amongst AMD and HMC sufferers than amongst people within the NC group. Particularly, MCP-1 concentrations had been 1154.0 ± 158.2 pg/mL within the AMD group, 1385.2 ± 158.2 pg/mL within the HMC group; each of them had been increased than the focus (741.3 ± 173.4 pg/mL) within the management group. We then carried out receiver working attribute curve evaluation to guage the diagnostic energy of the molecular detection know-how and decide the optimum cut-off level to tell apart between NC and blinding eye illness teams. The world underneath the curve (AUC) worth for the NC and blinding eye illness teams was 0.97 (Fig. 7D), thus the approach was in a position to precisely establish AMD and HMC sufferers from the members with a sensitivity of 95.0% and specificity of 80.0%. Moreover, a threshold worth of 1282 pg/mL was additional discovered to be efficient in differentiating AMD and HMC sufferers with a specificity of 86.0% (Fig. 7E). These outcomes point out that this know-how has the potential to be utilized within the prognosis and differentiation of AMD and HMC. Though additional multicenter scientific trials and extra experimental knowledge are essential to validate these findings, our developed molecular detection know-how supplies a promising resolution to the present limitations in widespread screening and early discovery of blinding eye ailments.
Molecular detection know-how for blinding eye ailments. (A) Schematic illustration of pattern acquisition, preprocessing, and evaluation utilizing the developed molecular detection know-how. (B) Correlation coefficients between detection outcomes obtained by ELISA and people obtained utilizing the developed know-how amongst sufferers with blinding eye ailments. (C) MCP-1 concentrations in AH samples from NC, and AMD, and HMC sufferers, as measured utilizing our developed detection know-how. (D) Receiver working attribute curves demonstrating the diagnostic efficiency of MCP-1 concentrations in blinding eye illness sufferers. (E) Receiver working attribute curve of MCP-1 to discriminate HMC from AMD sufferers